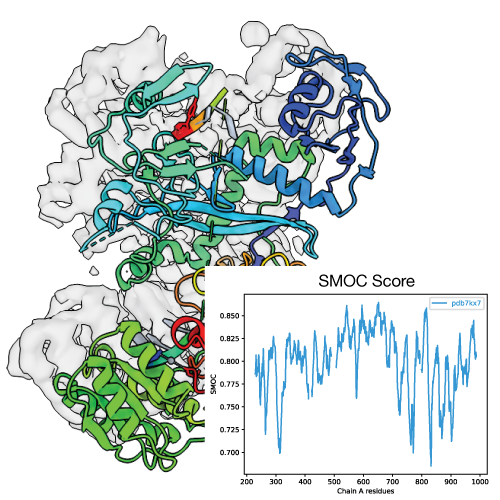
TEMPy2 offers a variety of local and global validation scores such as LoQFit, SMOC, SCCC and more.
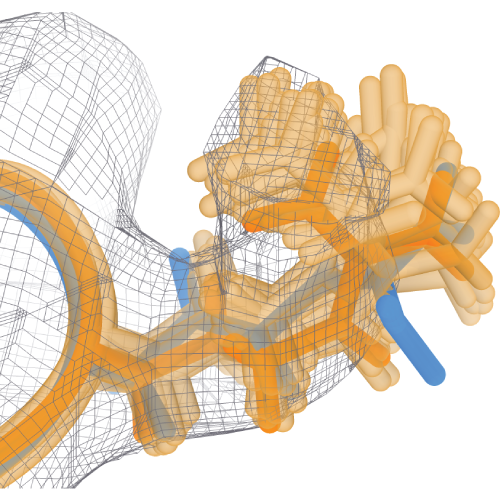
TEMPy-REFF offers B-factor refinement and can produce ensembles which better explain underlying dynamics.
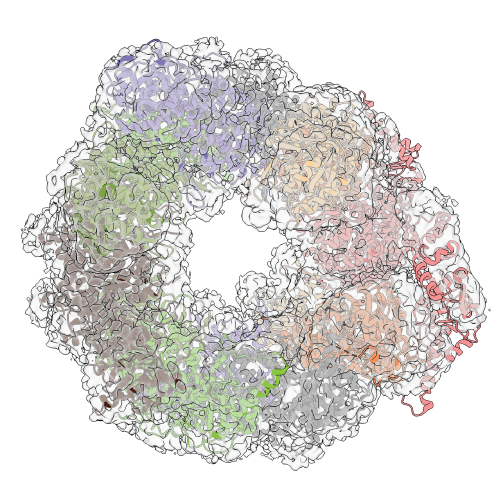
Build models into low resolution density with γ-TEMPy using a genetic algorithm.
TEMPy is used in the following projects and papers